In 1984, Greenberg at Harvard University in Boston discovered that serum addition to quiescent cells in culture could rapidly stimulate c-fos gene transcription (1). The induction reached maximum within 15 min and the c-fos mRNA level peaked at about 30 min after serum stimulation. This phenomenon led to a series of important discoveries in the area of gene transcription regulation. c-fos is a cellular homolog of the FBJ murine osteosarcoma virus transforming gene v-fos. Since it is rapidly activated without a need of new protein synthesis, c-fos was classified as an immediate early gene. Its activation is required for cells, which have exited the cell cycle, to reenter G1 phase of cell cycle and subsequently enter mitosis. Later, it was discovered that in addition to serum, growth factors and other mitogens could activate c-fos (2). Subsequent research has focused on the signaling upstream of c-fos gene and was aimed to explain why serum could activate c-fos. Several DNA elements have been identified in the c-fos promoter region (3-6), but a particular attention was given to a short sequence located about 300bp upstream of the transcription initiation start site. Treisman named this sequence Serum Response Element (SRE). It is an A/T rich core flanked by an inverted repeat, CCATATTAGG. This element was necessary and sufficient to render a heterologous promoter responsive to serum. Comparison of this element with the cytoskeletal actin gene promoter revealed a similar sequence (7). To date, about 30 genes have been identified that contain CC(A/T)6GG sequence in their promoter regions (8, 9) (Table 1). These genes include many immediate early genes such as c-fos, fosB, junB, egy-1, egy-2, etc. neuronal genes such as nur77, nurr1, etc. and muscle genes such as skeletal alpha-actin, alpha-myosin heavy chain (alpha-MHC), ß-myosin heavy chain (ß-MHC), myosin light chain (MLC), SM22alpha, telokin, troponin, tropomyosin, calponin, atrial natriuretic factor (ANF), Sarcoplasmic reticulum Ca2+-ATPase (SERCA), dystrophin, creatine kinase M, etc. Because of its special DNA sequence structure, SRE element is also referred to as CArG box. Treisman also identified the transcription factor, protein that binds to this element to control particular gene expression, and he named this protein Serum Response Factor (SRF).
What is SRF?
In humans, SRF gene is located on chromosome 6p21.1, and is 10607 base pair (bp) long. It contains 7 exons. The full length of its mRNA is 4201bp, exon1 (1-871), exon2 (872-1138), exon3 (1139-1400), exon4 (1401-1520), exon5 (1521-1712), exon6 (1713-1789), exon7 (1790-4201) (Fig. 1). The coding region is from 359 to 1885. By alternative splicing, several RNA transcripts could be generated. Conventionally, two mRNA species, 4.5kb and 2.5kb have been distinguished by Northern hybridization. In mouse, it has been reported that four RNA isoforms could be identified, depending on tissue type (10). Among these four isoforms, SRF-L contained all 7 exons, corresponding to the 4.5kb species in human. SRF-M isoform was lacking exon5 and functioned as a dominant negative mutant, which repressed SRF-dependent transcription (11). SRF-S lacking both exon5 and exon4 was only identified in the aorta. SRF-I isoform was the shortest one, contained only exons1, 2, 6 and 7, and was only detected in embryonic tissues.
| Table 1. Genes with SRF binding sites | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
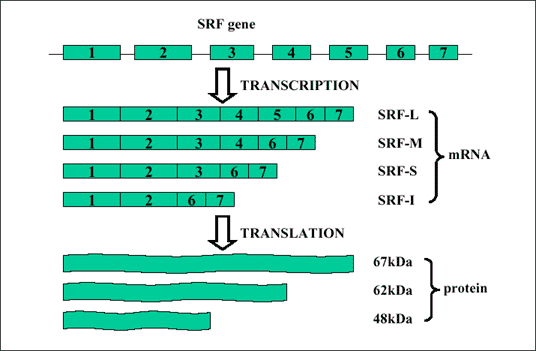 |
| Fig .1. Diagrammatic presentation of SRF gene structure and alternative splicing. SRF gene contains seven exons. Four mRNA isoforms have been identified by RT-PCR and three protein species have been identified by Western blotting. |
SRF protein contains 508 amino acids and is visualized as a 67kDa band on Western blot. It contains three major domains: (1) a SRE DNA binding domain; (2) a transactivation domain; and (3) several phosphorylation sites. Deletion analysis identified the DNA binding and dimerization domain to the region between amino acids 133 and 222 (12). This 90-amino-acid core domain is sufficient for DNA binding, dimerization and interaction with the accessory factors. This domain is highly conserved among eukaryotes. Even in those evolutionarily divergent species such as yeasts and plants an extensive homology has been recognized. This region is designated the MADS box, standing for MCM-1 from yeasts (13), Agamous and Deficiens from plants (14), and SRF from animals. Even between fruit fly and human, there is a 93% homology in SRF (15). The transcription activation domain is located in the C-terminal region of the SRF protein, within amino acids 339 to 508 region (16).
Biological Functions of SRF
SRF is an important regulator of numerous genes associated with cell growth and differentiation. SRF also regulates transcription resulting from treatment of cells with neurotrophins (17), neurotransmitters and agents that raise intracellular calcium levels (18, 19), stress agents, and viral activators (20, 21). Therefore, identification of the mechanism by which SRF mediates the activation of genes and regulation of the SRF gene itself is important for understanding these processes.
Inhibition of SRF by microinjection of anti-SRF antibodies or the expression of antisense SRF RNA suppressed muscle marker gene expression and blocked the differentiation of myoblasts to myotubes (22, 23). Moreover, SRF gene knockout demonstrated that homozygous SRF-/- mouse embryos failed to develop mesoderm (9). Mouse embryos lacking SRF developed normally until day E6.5. However, the mesodermal germ layer did not form and as a result, SRF-negative embryos died in utero between age E8.5 and E12.5. The fact that srf-/- embryos developed normally up to E6.5 and continued to grow even in the absence of mesoderm suggests that SRF is not a condition sine qua non for normal cell proliferation. In vitro study also supports this notion (24). By specific cardiac SRF transgenesis, we found that overexpression of SRF caused hypertrophic cardiomyopathy in mouse and the mouse died of heart failure within 6 months after birth (25). Overexpression of a mutant SRF that has no DNA binding activity severely damaged either embryogenesis or early development of the mouse, depending on the transgene copy number (26). On the other hand, overexpression of antisense SRF gave the mouse a better cardiac performance (Chai et al., unpublished data). All these transgenic data suggest that sufficient SRF is needed for embryogenesis and early development. For adults, maintaining relatively low level of SRF is more beneficial (Fig. 2).
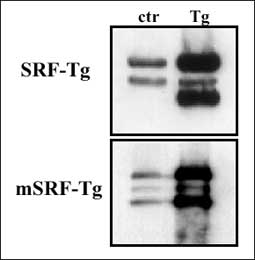 |
Fig. 2. Western blots showing overexpression of SRF (upper) and SRF mutant (mSRF) (lower) in transgenic mice compared to normal ones (25, 26). |
Studies performed on transgenic mice with different CArG boxes with flanking sequences linked to lacZ (27) suggest that sequences immediately surrounding the CArG box determine the expression pattern and that CArG boxes with muscle specificity bind SRF with reduced affinity compared with those with direct ubiquitous expression.
Activation and Regulation of SRF
SRF can be activated by a variety of agents, including serum, lysophosphatidic acid (LPA), anisomysin, mitogens, lipopolysaccharide (LPS), 12-O-tetradecanoylphorbol-13-acetate (TPA), redox, cytokines, tumor necrosis factor-a (TNFa); agents that elevate intracellular calcium levels; viral activator proteins such as the human T-cell lymphotropic virus type-1 activator protein Tax-1 and the hepatitis B virus activator protein pX; activated oncogenes including v-src, v-fps, v-ras, and the activated proto-oncogene c-raf as well as extracellular stimuli such as antioxidants, UV light, and microgravity. SRF is regulated both by cellular signal transduction pathways and by its interaction with other transcription factors including Sp1, ATF6, GATA4, Nkx2.5, and myogenic regulatory factors. Several mechanisms have been shown to regulate SRF activity, including (a) association with positive and negative cofactors (28), (b) phosphorylation-dependent changes in DNA binding (29), (c) alternative RNA splicing (10, 11), and regulated nuclear translocation (30).
There are two general classes of signaling mechanisms involving the SRF regulating SRE activity. A ternary complex factors (TCF)-dependent pathway involves the ras-raf-MAPK-ERK cascade (31). Both phosphorylation of TCFs and the binding of TCFs to SRF are required for activation of the SRE by this pathway. A TCF-independent pathway involves the Rho family of GTPases (32). Several MAPK pathways have been found to activate TCF. Among them, the extracellular-signal regulated kinase (ERK) can be activated by the extracellular stimulations mentioned above to phosphorylate TCF via Ras-Raf-MEK1/MEK2. The Jun N-terminal kinases (JNKs), also known as stress-activated protein kinases (SAPKs), can be turned on by cellular stress such as heat or UV light through small GTPases such as Rac and Cdc42 to phosphorylate TCF. Another subfamily of MAPK, p38/Mpk2/RK, could be also involved in the TCF activation. On the other hand, the biochemistry of SRF activation and the signaling pathways still remain unclear. It was recognized that all the factors that stimulate SRF expression have ability to activate the small GTPase RhoA. Therefore, it has been proposed that this might be the path how SRF is triggered. Ca2+/calmodulin-dependent kinase (Cam) might be involved in the process of SRF activation by cytoplasmic Ca2+ elevation (Fig.3).
Potential Implications for Tissue Injury Healing
As described above, genes under SRF regulation are mainly classified into three groups: immediate early genes, muscle genes and neuronal genes. Many of them are involved in cellular response to growth factor stimulation and tissue injury. Since SRF is an important regulator of these genes involved in cell growth and differentiation one of its important functions may be involvement in tissue injury and gastrointestinal ulcer healing. Since gastrointestinal injury is usually associated with the destruction of epithelium, smooth muscle structures and neural network and the healing process requires activation of numerous growth factors, it is reasonable to speculate that SRF might play an important role in gastrointestinal recovery from various injuries. Healing of these injuries requires regeneration of epithelial, muscle and neural structures and the neural/brain gut interactions play important role in gastrointestinal mucosal and pancreatic function, defense repair and healing (63-68).
Support: This work was supported by the VA Medical Research Service, REAP and VA Merit Review Awards A.S.T. Dr J. Chai is an Associate Investigator in the Research Enhancement Award Program at the VA Medical Center in Long Beach.
- Greenberg ME, Ziff EB. Stimulation of 3T3 cells induces transcription of the c-fos proto-oncogene. Nature. 1984; 311:433-8.
- Rollins BJ, Stiles CD. Serum-inducible genes. Adv Cancer Res. 1989;53:1-32.
- Tresman R. Identification of a protein-binding site that mediates transcriptional response of the c-fos gene to serum factors. Cell. 1986;46:567-577.
- Gilman MZ, Wilson RN, Weinberg RA. Multiple protein-binding sites in the 5-flanking region regulate c-fos expression. Mol Cell Biol. 198;6:4305-4316.
- Prywes R, Roeder RG. Inducible binding of a factor to the c-fos enhancer. Cell. 1986; 47: 777-84.
- Greenberg ME, Siegfried Z, Ziff EB. Mutation of the c-fos gene dyad symmetry element inhibits serum inducibility of transcription in vivo and the nuclear regulatory factor binding in vitro. Mol Cell Biol. 1987; 7:1217-1225.
- Mohun T, Garrett N, Treisman R. Xenopus cytoskeletal actin and human c-fos gene promoters share a conserved protein-binding site. EMBO J. 1987; 6:667-673.
- Cahill MA, Janknecht R. and Nordheim A. Signal uptake by the c-fos serum response element. Inducible Gene Expression. 1995; 2:39-72.
- Arsenian S, Weinhold B, Oelgeschlager M, Ruther U, Nordheim A. Serum response factor is essential for mesoderm formation during mouse embryogenesis. EMBO J. 1998; 17:6289-6299.
- Kemp PR, Metcalfe JC. Four isoforms of serum response factor that increase or inhibit smooth-muscle-specific promoter activity. Biochem J. 2000; 345:445-451.
- Belaguli NS, Zhou W, Trinh TH, Majesky MW, Schwartz RJ. Dominant negative murine serum response factor: alternative splicing within the activation domain inhibits transactivation of serum response factor binding targets. Mol Cell Biol. 1999;19: 4582-4591.
- Norman C, Runswick M, Pollock R, Treisman R. Isolation and properties of cDNA clones encoding SRF, a transcription factor that binds to the c-fos serum response element. Cell. 1988;55:989-1003.
- Shre P, Sharrocks AD. The ETS-domain transcription factors Elk-1 and SAP-1 exhibit differential DNA binding specificities. Nucleic Acids Res. 1995;23:4698-4706.
- Sommer H, Beltran JP, Huijser P, Pape H, Lonnig WE, Saedler H, Schwarz-Sommer Z. Deficiens, a homeotic gene involved in the control of flower morphogenesis in Antirrhinum majus: the protein shows homology to transcription factors. EMBO J. 1990; 9:605-613.
- Affolter M, Montagne J, Walldorf U, Groppe J, Kloter U, LaRosa M, Gehring WJ. The Drosophila SRF homolog is expressed in a subset of tracheal cells and maps within a genomic region required for tracheal development. Development. 1994;120: 743-753.
- Johansen FE, Prywes R. Identification of transcriptional activation and inhibitory domains in serum response factor (SRF) by using GAL4-SRF constructs. Mol Cell Biol. 1993;13:4640-4647.
- Sheng M, Dougan ST, McFadden G, Greenberg ME. Calcium and growth factor pathways of c-fos transcriptional activation require distinct upstream regulatory sequences. Mol Cell Biol. 1988; 8:2787-2796.
- McDonough PM, Hanford DS, Sprenkle AB, Mellon NR, Glembotski CC. Collaborative roles for c-Jun N-terminal kinase, c-Jun, serum response factor, and Sp1 in calcium-regulated myocardial gene expression. J Biol Chem. 1997; 272:24046-24053.
- Misra RP, Bonni A, Miranti CK, Rivera VM, Sheng M, Greenberg ME. L-type voltage-sensitive calcium channel activation stimulates gene expression by a serum response factor-dependent pathway. J Biol Chem. 1994;269:25483-25493.
- Avantaggiati ML, Natoli G, Balsano C, Chirillo P, Artini M, De Marzio E, Collepardo D, Levrero M. The hepatitis B virus (HBV) pX transactivates the c-fos promoter through multiple cis-acting elements. Oncogene. 1993;8:1567-1574.
- Fujii M, Tsuchiya H, Chuhjo T, Minamino T, Miyamoto K, Seiki M. Serum response factor has functional roles both in indirect binding to the CArG box and in the transcriptional activation function of human T-cell leukemia virus type I Tax. J Virol. 1994; 68:7275-7283.
- Soulez M, Rouviere CG, Chafey P, Hentzen D, Vandromme M, Lautredou N, Lamb N, Kahn A, Tuil D. Growth and differentiation of C2 myogenic cells are dependent on serum response factor. Mol Cell Biol. 1996;16:6065-6074.
- Gauthier-Rouviere C, Vandromme M, Tuil D, Lautredou N, Morris M, Soulez M, Kahn A, Fernandez A, Lamb N. Expression and activity of serum response factor is required for expression of the muscle-determining factor MyoD in both dividing and differentiating mouse C2C12 myoblasts. Mol Biol Cell. 1996; 7:719-729.
- Schratt G, Weinhold B, Lundberg AS, Schuck S, Berger J, Schwarz H, Weinberg RA, Ruther U, Nordheim A. Serum response factor is required for immediate-early gene activation yet is dispensable for proliferation of embryonic stem cells. Mol Cell Biol. 2001;21:2933-2943.
- Zhang X, Azhar G, Chai J, Sheridan P, Nagano K, Brown T, Yang J, Khrapko K, Borras AM, Lawitts J, Misra RP, Wei JY. Cardiomyopathy in transgenic mice with cardiac-specific overexpression of serum response factor. Am J Physiol Heart Circ Physiol. 2001;280:H1782-1792.
- Zhang X, Chai J, Azhar G, Sheridan P, Borras AM, Furr MC, Khrapko K, Lawitts J, Misra RP, Wei JY. Early postnatal cardiac changes and premature death in transgenic mice overexpressing a mutant form of serum response factor. J Biol Chem. 2001; 276:40033-40040.
- Chang PS, Li L, McAnally J, Olson EN. Muscle specificity encoded by specific serum response factor-binding sites. J Biol Chem. 2001276:17206-17212.
- Treisman R. Ternary complex factors: growth factor regulated transcriptional activators. Curr Opin Genet Dev. 1994; 4:96-101.
- Manak JR, Prywes R. Mutation of serum response factor phosphorylation sites and the mechanism by which its DNA-binding activity is increased by casein kinase II. Mol Cell Biol. 1991;11:3652-3659.
- Camoretti-Mercado B, Liu HW, Halayko AJ, Forsythe SM, Kyle JW, Li B, Fu Y, McConville J, Kogut P, Vieira JE, Patel NM, Hershenson MB, Fuchs E, Sinha S, Miano JM, Parmacek MS, Burkhardt JK, Solway J. Physiological control of smooth muscle-specific gene expression through regulated nuclear translocation of serum response factor. J Biol Chem. 2000; 275:30387-30393.
- Johansen FE, Prywes R. Two pathways for serum regulation of the c-fos serum response element require specific sequence elements and a minimal domain of serum response factor. Mol Cell Biol. 1994; 14: 5920-5928.
- Hill CS, Wynne J, Treisman R. The Rho family GTPases RhoA, Rac1, and CDC42Hs regulate transcriptional activation by SRF. Cell. 1995;1:1159-1170.
- Lazo PS, Dorfman K, Noguchi T, Mattei MG, Bravo R. Structure and mapping of the fosB gene. fosB downregulates the activity of the fosB promoter. Nucleic Acids Res. 1992; 20:343-350.
- Perez-Albuerne ED, Schatteman G, Sanders LK, Nathans D. Transcriptional regulatory elements downstream of the junB gene. Proc Natl Acad Sci U S A. 1993; 90:11960-11964.
- Wu BJ, Williams GT, Morimoto RI. Detection of three protein binding sites in the serum-regulated promoter of the human gene encoding the 70-kDa heat shock protein. Proc Natl Acad Sci U S A. 1987;84:2203-2207.
- Qureshi SA, Rim M, Bruder J, Kolch W, Rapp U, Sukhatme VP, Foster DA. An inhibitory mutant of c-Raf-1 blocks v-Src-induced activation of the egy-1 promoter. J Biol Chem. 1991; 266:20594-20597.
- Gius D, Cao XM, Rauscher FJ 3rd, Cohen DR, Curran T, Sukhatme VP. Transcriptional activation and repression by Fos are independent functions: the C terminus represses immediate-early gene expression via CArG elements. Mol Cell Biol. 1990; 10:4243-4255.
- Latinkic BV, OBrien TP, Lau LF. Promoter function and structure of the growth factor-inducible immediate early gene cyr61. Nucleic Acids Res. 1991; 19:3261-3267.
- Chung KC, Gomes I, Wang D, Lau LF, Rosner MR. Raf and fibroblast growth factor phosphorylate Elk1 and activate the serum response element of the immediate early gene pip92 by mitogen-activated protein kinase-independent as well as -dependent signaling pathways. Mol Cell Biol. 1998; 18:2272-2281.
- Ng SY, Gunning P, Liu SH, Leavitt J, Kedes L. Regulation of the human beta-actin promoter by upstream and intron domains. Nucleic Acids Res. 1989; 17:601-615.
- Framson P, Bornstein P. A serum response element and a binding site for NF-Y mediate the serum response of the human thrombospondin 1 gene. J Biol Chem. 1993; 268:4989-4996.
- Moiseyeva EP, Weller PA, Zhidkova NI, Corben EB, Patel B, Jasinska I, Koteliansky VE, Critchley DR. Organization of the human gene encoding the cytoskeletal protein vinculin and the sequence of the vinculin promoter. J Biol Chem. 1993; 268:4318-4325.
- Tan TH, Huang GP, Sica A, Ghosh P, Young HA, Longo DL, Rice NR. Kappa B site-dependent activation of the interleukin-2 receptor alpha-chain gene promoter by human c-Rel. Mol Cell Biol. 1992; 12:4067-4075.
- Castillo SO, Xiao Q, Lyu MS, Kozak CA, Nikodem VM. Organization, sequence, chromosomal localization, and promoter identification of the mouse orphan nuclear receptor Nurr1 gene. Genomics. 1997; 41:250-257.
- Williams GT, Lau LF. Activation of the inducible orphan receptor gene nur77 by serum growth factors: dissociation of immediate-early and delayed-early responses. Mol Cell Biol. 1993; 13:6124-6136. Miwa T, Kedes L. Duplicated CArG box domains have positive and mutually dependent regulatory roles in expression of the human alpha-cardiac actin gene. Mol Cell Biol. 1987; 7:2803-2813.
- Muscat GE, Gustafson TA, Kedes L. A common factor regulates skeletal and cardiac alpha-actin gene transcription in muscle. Mol Cell Biol. 1988; 8:4120-4133.
- Carson JA, Fillmore RA, Schwartz RJ, Zimmer WE. The smooth muscle gamma-actin gene promoter is a molecular target for the mouse bagpipe homologue, mNkx3-1, and serum response factor. J Biol Chem. 2000; 275:39061-39072.
- Kim JH, Johansen FE, Robertson N, Catino JJ, Prywes R, Kumar CC. Suppression of Ras transformation by serum response factor. J Biol Chem. 1994; 269:13740-13743.
- Molkentin JD, Jobe SM, Markham BE. Alpha-myosin heavy chain gene regulation: delineation and characterization of the cardiac muscle-specific enhancer and muscle-specific promoter. J Mol Cell Cardiol. 1996; 28:1211-1225.
- Huang WY, Chen JJ, Shih N, Liew CC. Multiple muscle-specific regulatory elements are associated with a DNase I hypersensitive site of the cardiac beta-myosin heavy-chain gene. Biochem J. 1997; 327:507-512.
- Henderson SA, Spencer M, Sen A, Kumar C, Siddiqui MA, Chien KR. Structure, organization, and expression of the rat cardiac myosin light chain-2 gene. Identification of a 250-base pair fragment which confers cardiac-specific expression. J Biol Chem. 1989; 264:18142-18148.
- Katoh Y, Loukianov E, Kopras E, Zilberman A, Periasamy M. Identification of functional promoter elements in the rabbit smooth muscle myosin heavy chain gene. J Biol Chem. 1994; 269:30538-30545.
- Yamamura H, Masuda H, Ikeda W, Tokuyama T, Takagi M, Shibata N, Tatsuta M, Takahashi K. Structure and expression of the human SM22alpha gene, assignment of the gene to chromosome 11, and repression of the promoter activity by cytosine DNA methylation. J Biochem (Tokyo). 1997; 122:157-167.
- Herring BP, Smith AF. Telokin expression in A10 smooth muscle cells requires serum response factor. Am J Physiol. 1997; 272:C1394-1404.
- Wang G, Yeh HI, Lin JJ. Characterization of cis-regulating elements and trans-activating factors of the rat cardiac troponin T gene. J Biol Chem. 1994; 269:30595-30603.
- Toutant M, Gauthier-Rouviere C, Fiszman MY, Lemonnier M. Promoter elements and transcriptional control of the chicken tropomyosin gene. Nucleic Acids Res. 1994; 22:1838-1845.
- Miano JM, Carlson MJ, Spencer JA, Misra RP. Serum response factor-dependent regulation of the smooth muscle calponin gene. J Biol Chem. 2000;275:9814-9822.
- Sprenkle AB, Murray SF, Glembotski CC. Involvement of multiple cis elements in basal- and alpha-adrenergic agonist-inducible atrial natriuretic factor transcription. Roles for serum response elements and an SP-1-like element. Circ Res. 1995; 77:1060-1069.
- Baker DL, Dave V, Reed T, Misra S, Periasamy M. A novel E box/AT-rich element is required for muscle-specific expression of the sarcoplasmic reticulum Ca2+-ATPase (SERCA2) gene. Nucleic Acids Res. 1998; 26:1092-1098.
- Klamut HJ, Gangopadhyay SB, Worton RG, Ray PN. Molecular and functional analysis of the muscle-specific promoter region of the Duchenne muscular dystrophy gene. Mol Cell Biol. 1990; 10:193-205.
- Vincent CK, Gualberto A, Patel CV, Walsh K. Different regulatory sequences control creatine kinase-M gene expression in directly injected skeletal and cardiac muscle. Mol Cell Biol. 1993; 13:1264-1272.
- Warzecha Z, Dembinski A, Ceranowicz P, Stachura J, Tomaszewska R, Konturek SJ. Effect of sensory nerves and CGRP on the development of caerulein-induced pancreatitis and pancreatic recovery. J Physiol Pharmacol. 2001; 52:679-704.
- Kwiecien S, Brzozowski T, Konturek PC, Konturek SJ, Pawlik M, Pajdo R, Drozdowicz D, Ptak A, Hahn EG. Effect of central and peripheral actions of histamine and its metabolite N-alpha methyl histamine on gastric secretion and acute gastric lesions. J Physiol Pharmacol. 2001; 52:625-638.
- Konturek PC, Brzozowski T, Meixner H, Ptak A, Hahn EG, Konturek SJ. Central and peripheral neural aspects of gastroprotective and ulcer healing effects of lipopolysaccharides. J Physiol Pharmacol. 2001;52:611-623.
- Brzozowski T, Konturek PC, Pajdo R, Kwiecien S, Ptak A, Sliwowski Z, Drozdowicz D, Pawlik M, Konturek SJ, Hahn EG. Brain-gut axis in gastroproctection by leptin and cholecystokinin against ischemia-reperfusion induced gastric lesions. J Physiol Pharmacol. 2001; 52:583-602.
- Pawlik M, Ptak A, Pajdo r, Konturek PC, Brzozowski t, Konturek SJ. Sensory nerves and calcitonin gene related peptide in the effect of ischemic preconditioning on acute and chronic gastric lesions induced by ischemia-reperfusion. J Physiol Pharmacol. 2001; 52:569-581.
- Warzecha Z, Dembinski A, Brzozowski T, Ceranowicz P, Dembinski M, Stachura J, Konturek SJ. Histamine in stress ulcer prophylaxis in rats. J Physiol Pharmacol. 2001; 52:407-421.