ONE SESSION OF EXERCISE REGULATES CATHEPSIN B ACTIVITY
IN THE LIVERS OF TRAINED AND UNTRAINED RATS
INTRODUCTION
Physical exercise causes adaptive changes, mainly in muscles (1) but it also influences many other organs and tissues in the organism, including liver, which serves as a very important energy source during exercise. Most of exercise-induced adaptive changes are beneficial (2). However, it has also been shown in human studies that strenuous exercise is a strong stressor and could result in splanchnic hypoperfusion with subsequent disturbances in liver homeostasis and energy, which can lead to liver damage (3, 4).
Cathepsin B is a lysosomal cysteine protease expressed constitutively. Cathepsin B is linked to general protein turnover and extracellular matrix (ECM) degradation. It functions as an exopeptidase at acidic pH, and as an endopeptidase in neutral pH, and it may enhance the activity of other proteases. It is also involved in autophagy and controls the number of lysosomes and autophagosomes in the cell (5-7). Autophagy is an evolutionarily conserved lysosomal pathway for degrading intracellular components, including cell organelles and cytoplasm. In normal cells under steady-state conditions, autophagy operates as a quality control mechanism, but it also plays an important role in response to a variety of homeostatic perturbations. It regulates energy generation by digesting cytoplasmic proteins and organelles, which allows cells to survive through starvation and stress conditions. Autophagy is also a lysosome-dependent pathway that can cause type II programmed cell death. Furthermore, lysosomal enzymes are engaged in necrosis (8, 9).
Very few studies have investigated the influence of exercise on cathepsin B generation and activity. It was shown that running increased cathepsin B levels in muscles and serum. In humans, changes in cathepsin B levels were correlated with fitness and memory function (10). In a study performed in horses, after 19 weeks of intense exercise, no increase in cathepsin B was observed in the cartilage of carpal joints (11).
This study aimed to investigate the influences of one session of exercise and endurance training on cathepsin B mRNA expression, protein levels, and enzyme activity in the livers of healthy rats.
MATERIALS AND METHODS
Exercise protocol
Sixty male Wistar rats were tested in this study. The rats had access to water ad libitum and Labofeed B. They were maintained in a 12 h/12 h light/dark cycle. All procedures in this study were approved by the Ethics Committee of the Medical University in Bialystok (Resolution No. 23/2011 on the proposal dated 27.04.2011). All protocols were performed according to EU regulations governing the treatment of laboratory animals. The experimental protocol was as previously described (12).
During the first 2 weeks of the experiment, rats were adapted to physical exercise. The adaptation consisted of 10 min of treadmill running every weekday, at 15 m/min. Then, rats were randomly assigned to two groups: untrained (UT, n = 30) and trained (T, n = 30). Training consisted of treadmill running, 5 days a week, for 6 weeks. During the first week, the running speed was constant (1200 m/h), but the exercise time was increased by 10 min each day, starting with 10 min of exercise per day. During week 2, rats were running 60 min per day, and the running speed was increased to 1500 m/h. During weeks 3 – 6, the running time was the same as during week 2, and the speed was increased to 1680 m/h. No additional stimulus was applied to enhance running.
The UT group remained sedentary during the entire training period. Twenty-four hours after the last training session, each group (UT and T) was randomly divided into three subgroups. In two groups (UTpre, n = 10 and Tpre, n = 10), the liver samples were collected under anesthesia (intraperitoneal chloral hydrate, 1 ml/100 mg body mass) and stored at –80°C for subsequent analysis. The remaining rats performed one session of exercise on a treadmill, running at 1680 m/h for 60 min. Liver samples were collected immediately after the exercise session (groups UT0h, n = 10 and T0h, n = 10) or 3 h after the acute exercise (UT3h, n = 10 and T3h, n = 10) and stored at –80°C for subsequent analyses. During this experiment, other samples were also collected (e.g., heart). All animals died as a consequence of sample collection. The scheme of experiment is presented in Fig. 1.
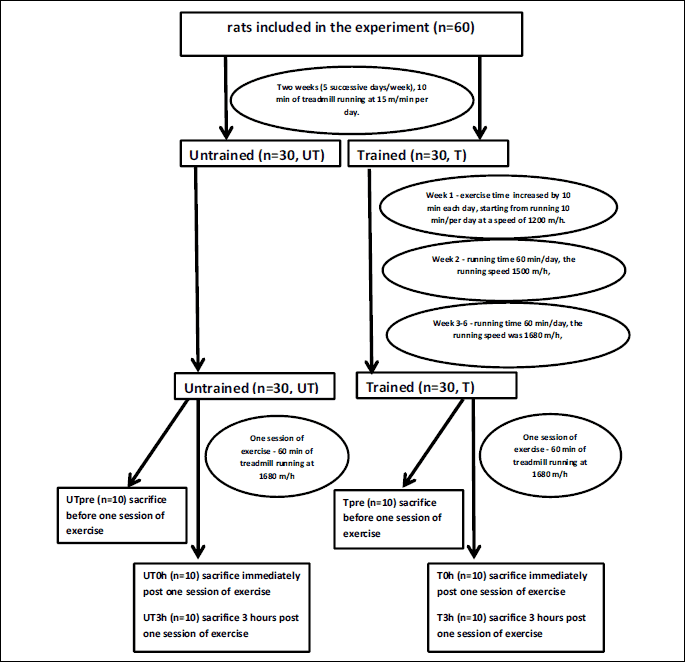
The mean body masses of rats on the day of acute exercise were 271 ± 11.6 g in the UT group and 283.17 ± 24.67 g in the T group.
Tissue homogenization
About 25 mg of rat liver tissue was placed in a round-bottom tube and homogenized with a steel ball (diameter 5 mm) in the TissueLyser (Qiagen, Germany). Homogenization was performed at room temperature at a frequency of 25 Hz for 5 min. The tissues intended for real-time PCR (RT-PCR) were homogenized in 750 µl of QIAzol Lysis Reagent (Qiagen, Germany). The tissues used to determine tissue protein levels were homogenized in 1 ml of 1 × PBS. The tissues intended for measuring the activity of cathepsin B was homogenized in 1 ml of deionized water. Samples intended for determining protein concentrations were subjected to two cycles of freezing (–20°C) and thawing (room temperature), then they were centrifuged at 5000 × g for 5 min at 4°C. All procedures were performed according to manufacturer instructions.
Isolation of total RNA
We isolated total RNA from rat liver tissue with a BioRobot EZ1 (Qiagen, Germany) and an EZ1 RNA Universal Tissue Kit (Qiagen, Germany), according to the manufacturer’s instructions. RNA quality and quantity were evaluated with a ND-1000 camera (NanoDrop Technologies, Wilmington, Delaware, USA). Then, total RNA samples were stored at –80°C.
Real time polymerase chain reaction
Real time polymerase chain reaction (RT-PCR) analyses of rat liver tissue sample was performed with an ABI 7500 thermocycler (Applied Biosystems, Waltham, Massachusetts, USA). To each well, we added the reaction mixture, which included the TaqMan RNA-to-CT 1-Step Kit (Applied Biosystems, Waltham, Massachusetts, USA) and primers, appropriately selected for the TaqMan Gene Expression Assay (Applied Biosystems, Waltham, Massachusetts, USA). Primers specifically targeted sequences in the cathepsin B gene and the control gene, glyceraldehyde 3-phosphate dehydrogenase (GAPDH). Next, we added total RNA to the reaction mixture in all wells, except the No Template Control (NTC), which received water instead of RNA (Sigma-Aldrich, Saint Louis, Missouri, USA). The PCR was performed according to the protocol recommended by Applied Biosystems. The test samples and control were assayed in duplicate, and the NTC was assayed in triplicate.
Gene expression levels were calculated with the comparative cycle threshold (CT) method. The CT of each sample was normalized to the CT of GAPDH as follows: ΔCT = CTresearch gene - CTGAPDH. The relative gene expression levels (ΔΔCT) were calculated by subtracting normalized CT values for the trained group from those of the untrained group, as follows: ΔΔCT = ΔCTtrained – ΔCTuntrained. Finally, the fold change in the mRNA level after treatment was calculated as: 2–ΔΔCT (13).
Measuring total protein levels
The total protein contents of rat liver tissue samples were measured with a colorimetric method (Pierce BCA Protein Assay Kit, Thermo Fisher Scientific, Waltham, Massachusetts, USA), according to the manufacturer’s instructions. Protein contents were measured at 562 nm on the BioTek Power Wave XS spectrophotometer (BioTek Instruments, Winooski, Vermont, USA).
Measuring cathepsin B concentrations
The concentration of cathepsin B in rat liver tissue samples was measured with the Rat Cathepsin B ELISA Kit (EIAab), according to the manufacturer’s instructions. Solutions were measured at 450 nm on the BioTek Power Wave XS spectrophotometer (BioTek Instruments, Winooski, Vermont, USA).
Measuring cathepsin B activity
Cathepsin B activity in rat liver tissue samples was measured with a spectrofluorimeter (LS-50B, PerkinElmer, USA). Fluorescence measurements were performed with induction at l = 355 nm and emission at λ = 460 nm. The substrate for cathepsin B was Z-Arg-Arg-AMC (N-CO2-L-arginylarginine-7-amino-4-methylocoumarin salt; Bachem, Biochemica GmbH, Heidelberg, Germany).
Statistical analysis
Results are reported as the median (minimum and maximum values). Two-way ANOVA was performed in order to analyze influence of both, training and time of sampling (i.e. pre exercise, at the completion of exercise, and 3 hours post exercise) on cathepsin B gene expression, protein concentration, and activity. Contrast analysis (on lsmeans) with Turkey method was performed regardless ANOVA results. P-values < 0.05 were considered statistically significant.
RESUTLS
One session of exercise did not influence cathepsin B gene expression or protein concentration at any investigated time point (Fig. 2 and Fig. 3, respectively). The effects of one session of exercise on cathepsin B activity are presented in Fig. 4. ANOVA revealed, that interactions between subgroup and training status were statistically significant (P = 0.003).
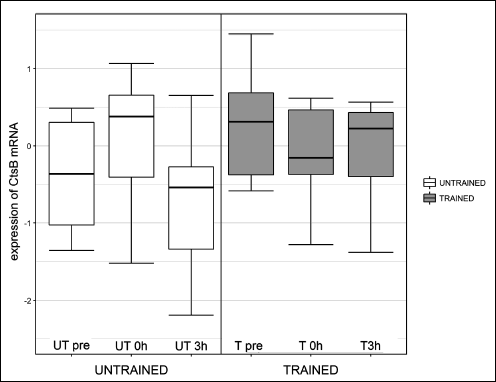 |
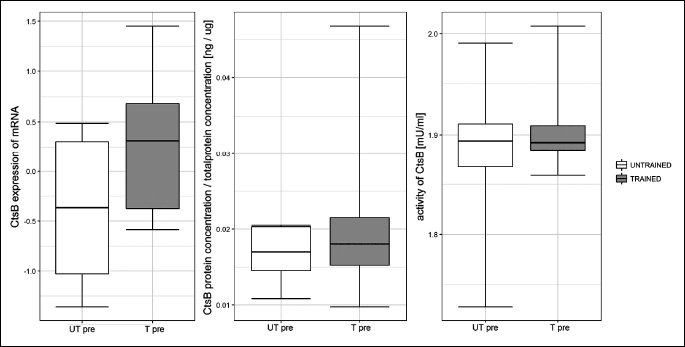
Fig. 2. Influence of one session of exercise on liver cathepsin B (Cts B) mRNA in healthy rats. One session of exercise did not influence cathepsin B gene expression at any investigated time point. |
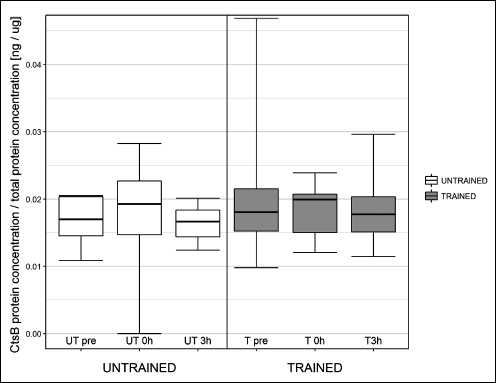 |
Fig. 3. Influence of one session of exercise on liver cathepsin B (Cts B) protein concentration in healthy rats. One session of exercise did not influence cathepsin B protein concentration at any investigated time point. |
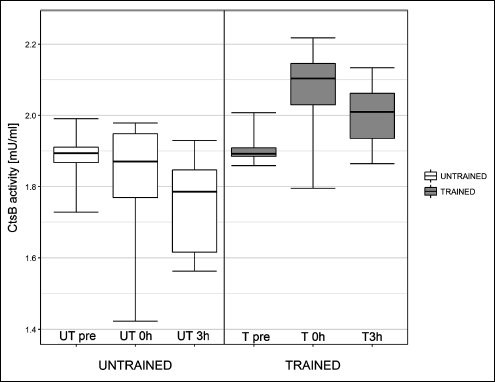 |
Fig. 4. Influence of one exercise session on liver cathepsin B (Cts B) activity in healthy rats. In untrained rats, cathepsin B activity was similar to basal levels (UT0h versus UTpre) immediately post exercise, and the activity declined at 3 h post exercise (UT3h versus UTpre; P = 0.027). In trained rats, cathepsin B activity increased immediately post exercise compared to basal levels (T0h versus Tpre: P = 0.005). |
In untrained rats, cathepsin B activity declined at UT3h compared to UTpre (P = 0.027). However, there was no difference in cathepsin B activity between UTpre and UT0h, and UT0h and UT3h. In trained rats, cathepsin B activity increased between Tpre and T0h (P = 0.005). There was no difference in cathepsin B activity between Tpre and T3h, and T0h and T3h.
Endurance training did not influence cathepsin B generation, either at the mRNA level or the protein level, or cathepsin B activity (Fig. 5).
In order to establish the influence of training and time at which samples were collected, on the activity of cathepsin B, a Bayesian hierarchical model was also employed. The ‘mean of posterior distribution’ (mean) value shows average of posterior distribution for a given parameter, while 95% HDI (highest density interval) low and high represent range in which 95% of posterior distribution density is located. If in case of investigated factor, both low and high end of 95% HDI is below, or above 0, it indicates a statistically significant influence of this factor on observed variability.
Training has a significant influence on cathepsin B activity, the time of sampling does not influence activity per se but interaction between training and sampling time does. Training did not influence basal activity od cathepsin B (UTpre versus Tpre, mean was –0.023 mU/ml, 95% HDI –0.0395 and 0.0897). In untrained animals activity od cathepsin B did not change immediately post exercise in compare to pre exercise level. The activity of cathepsin B dropped significantly 3 hours post exercise (UT3h versus UTpre, mean was 0.113 mU/ml, 95% HDI 0.011 – 0.223). There was no significant difference in cathepsin B activity between UT3h and UT0h.
In trained animals, activity of cathepsin B increased significantly in T0h in compare to Tpre (mean was –0.16 mU/ml, 95% HDI –0.249 and –0.0679). Cathepsin B activity remained higher that measured prior to exercise 3 hours after its cessation (mean was –0.093 mU/ml, 95% HDI –0.16 and –0.0172). Here the results obtained in Bayesian analysis differ from those obtained in ANOVA. There was no difference between T3h and T0h.
Bayesian analysis revealed significantly higher cathepsin B activity post one session of exercise in trained rats as compared with untrained (UT0h versus T0h, mean was 0.213mU/ml, 95% HDI 0.0885 – 0.339), and 3 hours post exercise (UT3h versus T3h, mean was 0.229 mU/ml, 95% HDI 0.119 – 0.341).
DISCUSSION
Our results showed that one session of exercise could regulate liver cathepsin B activity differently, depending on training status. However, 6-weeks of endurance training did not change basal liver cathepsin B activity. We hypothesized that cathepsin B activity changed in response to particular stress factors, which were absent in basal conditions, despite the training status.
Cathepsin B gene expression and protein concentrations were not influenced by either prolonged training or one session of exercise. Unaltered cathepsin B mRNA in the liver tissue after running was also revealed by Moon et al. (9). The main aim of this study was to examine the influence of exercise on cathepsin B generation and activity in the liver, however, because cathepsin B is involved in authophagy, we should admit here that the authophagic response to stress has two phases: first, a rapid increase in autophagy occurs within minutes or hours of exposure to stressful conditions, and it is entirely mediated by post-translational protein modifications; second, a delayed phase occurs that relies on the activation on specific transcription factors (9). The time points chosen in this study pertained to the early response. Besides, Bayod et al. (14) showed, that long-term moderate activity did not result in autophagic process in the liver tissue.
The increase in cathepsin B activity after one session of exercise in trained animals could contribute to homeostatic stability. This level of exercise required the consumption of a large amount of energy. In human studies, it was shown that 1 hour of cycling at 70% maximum work load capacity resulted in splanchnic hypoperfusion with subsequent liver damage, indicated by increased serum levels of liver fatty acid binding protein, alanine transaminase, aspartate transaminase, and alpha-glutathione S-transferase. In that type of setting, cathepsin B might regulate the process of autophagy to mediate a cytoprotective effect. Energy homeostasis might be maintained by the digestion of cytoplasmic proteins and organelles to generate energy substrates. Cathepsin B might also contribute to the removal of cell remnants and cytotoxic stimuli, and thus, control the adaptive response to stress (4, 9). Cathepsin B is also a negative feedback regulator of lysosomal biogenesis and autophagy; therefore, it could protect the cell from exaggerated autophagy, which could kill the cell (8). It should be mentioned here that cathepsin B participates in the pathophysiology of many diseases, such as rheumatoid arthritis, atherosclerosis, periodontitis, and pancreatitis. According to recent studies, it plays a crucial role in cancer development, by regulating tumor angiogenesis, invasion, and metastases; therefore, an increase in cathepsin B activity is potentially harmful (15). In our experiment, the increase in cathepsin B was significant, but transient. Three hours after the cessation of exercise, cathepsin B activity tended to decrease, compared to its activity immediately after exercise cessation.
It is difficult to interpret the decline in cathepsin B activity in untrained animals after one session of exercise. However, a previous study also observed that exercise had differential effects on the function of a particular organ, in trained and untrained individuals (12). In this study, the exercise session was very intense. It is possible that a delayed response to stress took place after the investigated time points. It is also possible that the stress was too intense for the untrained organism, and the response was inadequate. Here, we should mention that a decline in cathepsin B activity was associated with a dysfunction in autophagy, and this dysfunction was previously shown to be involved in the pathogenesis of nonalcoholic fatty liver disease (16, 17).
In conclusion, physical exercise influenced cathepsin B activity in the liver, and the type of influence depended on training status. Our results suggested that exercise intensity should be graded to match the training status. That is, training should start with low effort, and increasing effort should follow the increase in training status. However more study is required to support this statement, particularly in terms of the effects of exercise on the liver.
Conflict of interests: None declared.
REFERENCES
- Czarkowska-Paczek B, Zendzian-Piotrowska M, Bartlomiejczyk I, Przybylski J, Gorski J. The effect of acute and prolonged endurance exercise on transforming growth factor-beta1 generation in rat skeletal and heart muscle. J Physiol Pharmacol 2009; 60: 157-162.
- Kumral ZN, Sener G, Ozgur S, et al. Regular exercise alleviates renovascular hypertension-induced cardiac/endothelial dysfunction and oxidative injury in rats. J Physiol Pharmacol 2016; 67: 45-55.
- Tang Y, Li J, Gao C, et al. Hepatoprotective effect of quercetin on endoplasmic reticulum stress and inflammation after intense exercise in mice through phosphoinositide 3-kinase and nuclear-factor kappa-B. Oxid Med Cell Longev 2016; 2016: 8696587. doi: 10.1155/2016/8696587.
- van Wijck K, Lenaerts K, van Loon LJ, Peters WH, Buurman WA, Dejong CH. Exercise-induced splanchnic hypoperfusion results in gut dysfunction in healthy man. PLoS One 2011; 6: e22366. doi: 10.1371/journal.pone.0022366
- Aggarwal N, Sloane BF. Cathepsin B: Multiple roles in cancer. Proteomics Clin Appl 2014; 8: 427-427.
- Fais S. Cannibalism: a way to feed on metastatic tumors. Cancer Lett 2007; 258: 155-164.
- Man SM, Kanneganti TD. Regulation of lysosomal dynamics and authophagy by CTSB/cathepsin B. Authophagy 2016; 12: 2504-2505.
- Chang CP, Yang MC, Lei HY. Concavalin A/INF-gamma triggers authophagy-related necrotic hepatocyte death throu IRGM1-mediated lysosomal membrabe disruption. PLoS One 2011; 6: e28323. doi: 10.1371/journal.pone.0028323
- Pietrocola F, Izzo V, Niso-Santano M, et al. Regualtion of authophagy by stress-responsive transcription factors. Semin Cancer Biol 2016; 23: 310-322.
- Moon HY, Becke A, Berron D, et al. Running-induced systemic cathepsin B secretion is associated with memory function. Cell Metab 2016; 24: 332-340.
- Bowe FA, Murray RC, Jeffcott LB, Davies ME. Do the matrix degrading enzymes cathepsin B and D increase following a high intensity exercise regime? Osteoarthritis Cartilage 2007; 15: 343-349.
- Czarkowska-Paczek B, Zendzian-Piotrowska M, Gala K, Sobol M, Paczek L. Exercise differentially regulates renalase expression in skeletal muscle and kidney. Tohoku J Exp Med 2013; 231: 321-329.
- Livak KJ, Schmittgen TD Analysis of relative gene expression data using real-time quantitative PCR and the 2(-delta delta CT) method. Methods 2001; 25: 402-408.
- Bayod S, Del Valle J, Pelegri C, et al. Macroautophagic process was differentially modulated by long-term moderate exercise in rat brain and peripheral tissues. J Physiol Pharmacol 2014; 65: 229-239.
- Lei T, Ling X. IGF-1 promotes the growth and metastasis of hepatocellular carcinoma via the inhibition of proteasome-mediated cathepsin B degradation. World J Gastroenterol 2015; 21: 10137-10149.
- Fukuo Y, Yamashina S, Sonoue H, et al. Abnormality of autophagic function and cathepsin expression in the liver from patients with non-alcoholic fatty liver disease. Hepatol Res 2014; 44: 1026-1036.
- Inami Y, Yamashina S, Izumi K, et al. Hepatic steatosis inhibits autophagic proteolysis via impairment of autophagosomal acidification and cathepsin expression. Biochem Biophys Res Commun 2011; 412: 618-625.
A c c e p t e d : October 25, 2017